Jonas Koeppel
Group Leader (incoming)
ORCID: 0000-0003-1306-3994
EditEngineering structural variation to understand human genome function

Group Leader (incoming)
ORCID: 0000-0003-1306-3994
EditWhen do we truly understand a mammalian genome? Such understanding requires more than describing the function, or lack thereof, of each base pair. It also requires the ability to predict the consequences of sequence and structural variants and to design new genomes that function as intended. We still lack methods to systematically engineer genome architecture at larger scales through structural variation (SV). However, a flurry of new methods now puts “programmable SV engineering” within reach, making this an incredibly exciting time for this nascent field.
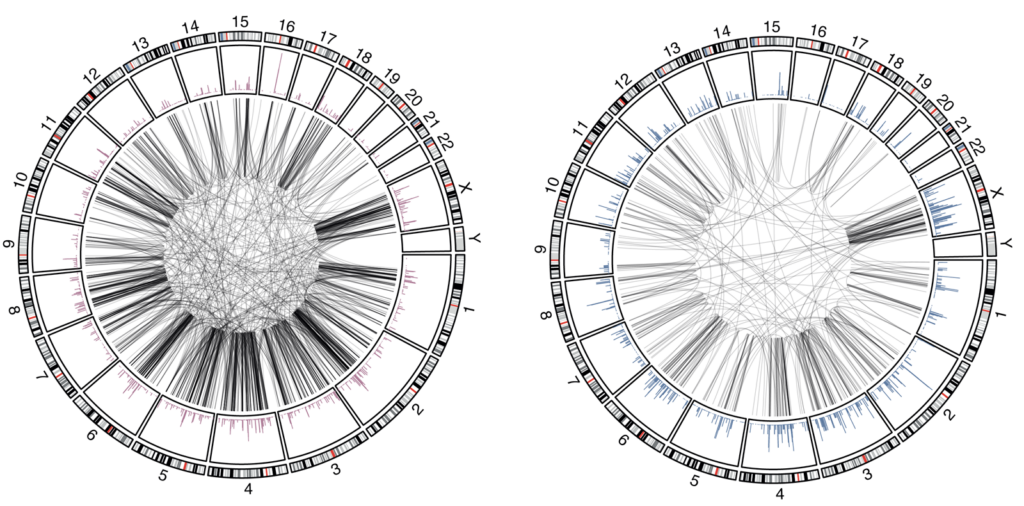
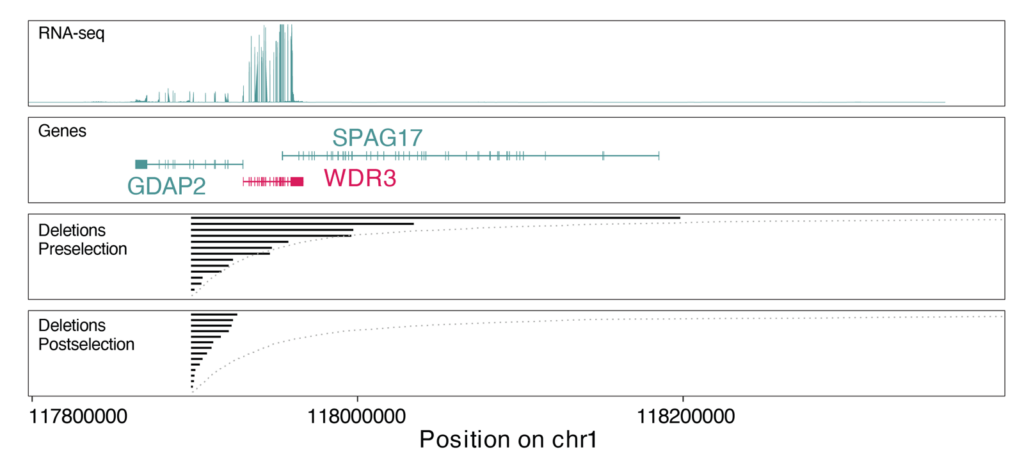
By combining complementary genome editing strategies, including prime editing, recombinases, and CRISPR-Cas3, we have generated highly engineered human cell systems that enable SV formation in mammalian genomes at an unprecedented scale. We pair these perturbation strategies with long-read sequencing, phage polymerase-based genotyping, and emerging single-cell approaches to directly resolve engineered variants and connect them to changes in fitness and gene expression. Together, these advances begin to make genome architecture experimentally tractable and provide a route to uncover design principles and essential elements embedded within the noncoding genome.
Our group will develop and apply scalable technologies to design, generate, and phenotype thousands of structural variants across mammalian genomes. We aim to map sequence dispensability genome-wide, uncover functional noncoding elements and architectural constraints, and generate datasets that support predictive and generative models of genome structure-function relationships. Our work will illuminate the mechanisms by which SVs cause disease and help define the genomic content required for mammalian cellular viability to establish principles for the design of minimal mammalian genomes.