Previous and current research
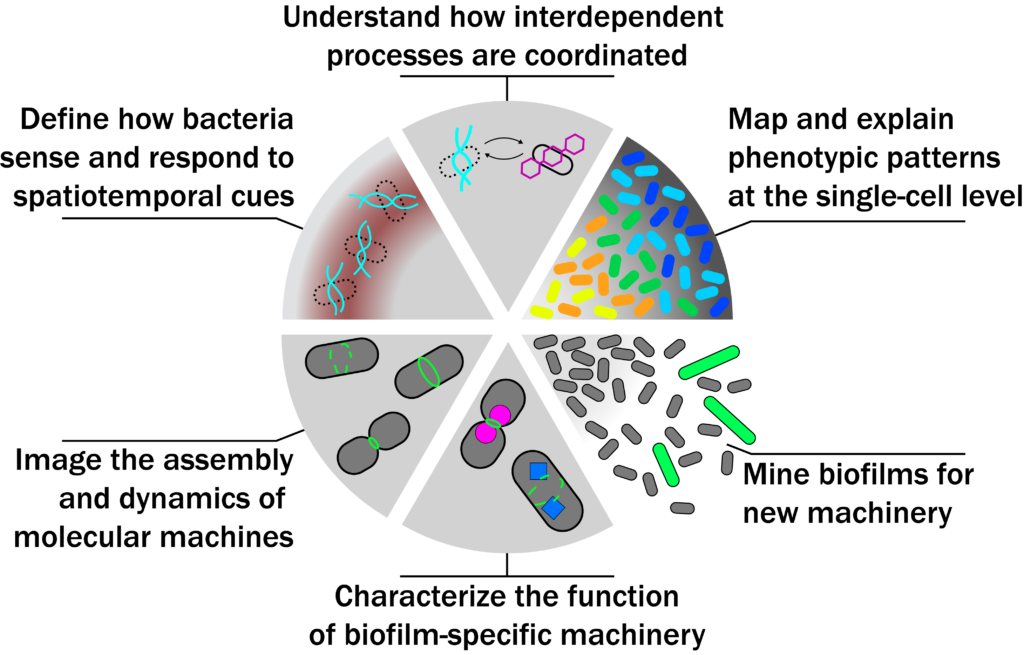
Most microbes on Earth live in dense multicellular communities called biofilms. Biofilms exist in every major habitat, shape earth’s climate, and cause devastating treatment-resistant infections. Bacterial biofilms carry out complex processes that individual bacteria do not: they become more virulent, stress-resistant, and antibiotic-tolerant; divide labour to increase efficiency; and develop via stereotyped changes in structure and function. How are these complex behaviours orchestrated by individual bacteria, and how can molecule- and cell-scale mechanisms explain biofilm-scale function? We focus on two research questions:
- How are complex, multicellular processes orchestrated by life’s simplest cells? Bacteria in biofilms become specialized, and this specialization is often patterned in space and time. We make 4D maps of phenotypic patterning, define molecular mechanisms by which bacteria sense and respond to spatiotemporal cues, and build models explaining how biofilm-scale function emerges from single-cell behavior.
- How do bacteria adapt their cell biology to life in biofilms? In biofilms, bacteria face steep nutrient gradients, mechanical confinement, and other stressors. In this environment, our understanding of bacterial cell biology breaks down, and antibiotics targeting this biology often fail. We investigate changes to the assembly and dynamics of essential molecular machines inside biofilm bacteria and use screening and -omics approaches to mine for biofilm-specific cell biology.
To learn more, see:
Future projects and goals
Topics in our group include:
- Biofilm matrix production: How is extracellular matrix production coordinated in biofilms? What are the molecular mechanisms by which specific matrix components are made by the right subpopulation, in the right place, and at the right time? How does this coordination lead to a functional and robust biofilm matrix?
- Cell division machinery in biofilms: How does the cell division complex, and especially the bacterial tubulin FtsZ, change its function in biofilms? How can we mine the biofilm to find new cell division proteins, especially tubulin binders?
- Biofilm imaging, analysis, and modelling: How can we improve our microfluidic and microscopy methods to see biofilms more clearly, especially at the subcellular level? How can machine learning help us extract meaning from large 4D data sets? How can modelling bridge the gap between molecular mechanisms and collective phenomena?
Our primary model is the opportunistic pathogen Pseudomonas aeruginosa, but our strain collection is growing!