Are protein domains indivisible?
Protein domains are considered to be indivisible units – but new research from EMBL-EBI shows that some can function even with big parts missing.
Researchers at EMBL-EBI have shown that protein domains, previously thought to be indivisible, complete units that make up proteins, can exist and function despite having lost large parts of their structure. Published in Genome Biology, the research into ‘domain atrophy’ offers new insights into the evolution, molecular mechanism, stability and function of protein domains.
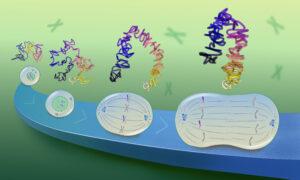
Proteins are everywhere, performing an endless variety of tasks to keep life moving along. They are made up of independent units called protein domains, which can be assembled in different combinations to build full proteins to order. Protein domains are defined by core structural elements, much in the same way a gear on a bike, watch or car can be defined by the number, size and arrangement of its cogs.
But are protein domains really indivisible? The researchers set out to answer this question computationally by searching and comparing millions of protein sequences and hundreds of protein structures in public databases. They identified a small percentage of protein domains that have been retained throughout evolution, functioning just like their ancestors, despite having undergone large-scale structural deletions.
“You can think of protein domains like cogs in an engine, with all these teeth that interlock and make each other move,” explains Ananth Prakash of EMBL-EBI. “You can assume that a cog will need all its teeth to make a good enough connection to work properly. But we found that with some protein domains, the engine parts keep moving even if a large number of those ‘teeth’ are missing.”
It is simply amazing that proteins can tolerate such extreme mutations.
“Normally, you’d expect large-scale deletions, which we call domain atrophy, to inhibit a domain’s function and cause unfolding,” adds Alex Bateman, head of Protein Sequence Resources at EMBL-EBI. “Given that a mutation in a single residue can lead to a human disease, it is simply amazing that proteins can tolerate such extreme mutations.”
The team started with two major public resources, UniProt and Pfam, selecting only the most robust entries and using a new algorithm to identify cases of partial protein domains. By painstakingly sifting through the results, like panning for gold, they collected a small set of protein domains with large-scale deletions in their core structural elements.
“Most atrophied domains retain structural elements or catalytic residues that are crucial for the normal functioning of a protein, which only makes sense,” says Ananth. “Interestingly, most of them appear to compensate for those atrophied sections by forming complexes with other partners.”
At the ISMB conference in 2014, members of the Bateman team at EMBL-EBI and the Pearson group at the University of Virginia discovered that they had both been working along the same lines. Both groups agree that domain atrophy is extremely rare and that for the most part, protein domains are indivisible, structural building blocks. But their findings reveal a fascinating phenomenon, and the shared terminologies developed by the groups will help future investigations into the molecular mechanisms that cause parts of these protein domains to disappear in the first place.
This post was originally published on EMBL-EBI News.
Related links
- Prakash A and Bateman A (2015). Domain atrophy creates rare cases of functional partial protein domains. Genome Biol. (in press); DOI: 10.1186/s13059-015-0655-8
- Triant D and Pearson W (2015). Most partial domains in proteins are alignment and annotation artifacts. Genome Biol. (in press); DOI: 10.1186/s13059-015-0656-7