No humans required
Match-making used to be a laborious task. People would fill in forms describing themselves and the kind of person they were looking for, and then someone would manually try to find a suitable date among other archived singletons. Today it’s all much quicker and easier: you just complete your profile online and computer algorithms do all the hard work of finding your dream partner.

Biologists looking for the partners that a given protein naturally has in the cell are on the edge of a similar technical revolution. There are many tools available for this task, but like the match-making methods of old they can be time-consuming and take a lot of effort. The new approach, which is reported today in Nature Biotechnology, provides an automated tool for scientists to track proteins and the molecules they interact with in living cells. Such studies provide clues about how the cell’s proteins connect with each other to carry out crucial functions like cell division. They can also shed light on what goes wrong in disease, and which proteins could be useful drug targets for pharmaceutical treatments.
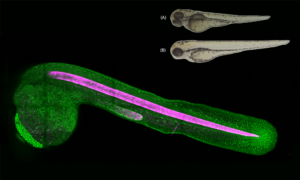
The new approach comes from work by Malte Wachsmuth, Christian Conrad and colleagues in the laboratory of Jan Ellenberg, Head of the Cell Biology and Biophysics Unit, who teamed up with Rainer Pepperkok and the Advanced Light Microscopy Facility at EMBL Heidelberg to develop new software that automates a powerful but previously slow technique called fluorescence correlation spectroscopy (FCS). In a standard FCS study, scientists label molecules of interest within the cell — specific proteins, for example — with a fluorescent tag. The cell is then placed under a microscope, and a laser is shone on the sample, causing the tag to emit light of a specific colour, such as green or red. Typically, the laser is focused on a tiny volume within the cell so that only a few light-emitting molecules are visible at any one time.
FCS allows scientists to see how quickly these molecules pass through this small volume, or whether one labelled molecule is usually present together with other proteins labelled with a different colour. From this data, they can work out the rough size of molecules, and which other proteins they bind to, among other properties. What’s made FCS slow in the past is that a human operator has had to identify a suitable cell to examine in detail, focus the microscope to zoom in on a small area, and then record a single-pixel movie — before repeating the process with another cell. After all this, the data have to be analysed mathematically to work out the properties of the molecules under study. “It’s very labour-intensive, and that’s a major issue”, says Wachsmuth.
It’s very labour-intensive, and that’s a major issue
The software developed by Wachsmuth and colleagues takes the human out of this equation and speeds up the process enormously. Whereas before a scientist could spend a whole day observing a single protein in a few cells, the automated process can analyse thousands of cells in the same time. To do so, cells are placed in a multi-well plate, with each well containing hundreds of cells laid out on a grid. The microscope peers down at a well, and the software identifies cells suitable for further analysis. It then automatically re-programs the microscope to zoom in a specific portion of the cell – a tiny part of the nucleus, for example. After recording what’s happening in this region, the microscope zooms out, looks for another cell, and starts over again.
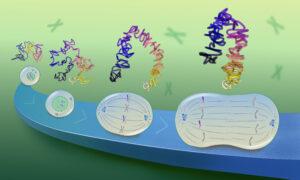
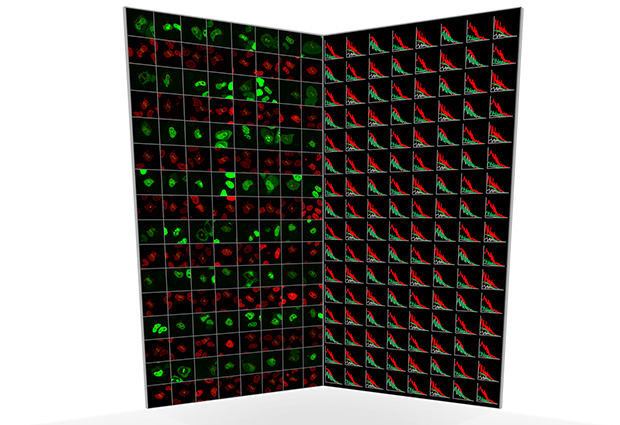
Not only does the software automate the collection of data, it also processes them to calculate the various properties of interest to researchers. “In the past, the analysis of FCS data has been as tedious as acquiring the data,” says Wachsmuth. To show that the new automated approach works, Wachsmuth and colleagues carried out two biological proof-of-principle studies. In one, they tagged chromatin, a protein component of chromosomes, with a red fluorescent tag, and then labelled 53 other proteins with green tags. By focusing on the nucleus, and looking for the co-movement of red and green proteins, the team were able to identify which of the proteins bound to chromatin.
In the second study, the team looked at two proteins that have been proposed to bind to each other during cell division. These proteins were labelled with red and green fluorescent tags, respectively, and then followed over the course of the cell cycle. “Nobody had ever shown that they really interact in a living cell, or when during the cell cycle they form a pair”, says Wachsmuth. By revealing when red and green proteins moved together (indicating that they are bound together) or on their own, the automated FCS approach revealed how the concentrations of these proteins fluctuated across the cell cycle, and when they came together to do their job.
This automated, high-throughput method will enable scientists to study many different proteins in a single experiment and see how they interact to form complex networks, which is essential for understanding how cells function. The software developed by Wachsmuth, Conrad and colleagues is freely available to download from the EMBL website, so researchers around the world will be able to take some of the hard work out of their FCS experiments