Rearranging chromosomes
Does rearranging chromosomes affect their function? EMBL scientists reveal uncoupling of 3D chromatin organisation and gene expression.

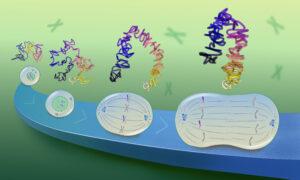
Molecular biologists long thought that domains in the genome’s 3D organisation control how genes are expressed. After studying highly rearranged chromosomes in fruit flies, EMBL researchers now reveal that while this is the case for some genes, their results challenge the generality of this for many others. Their results, published in Nature Genetics on 15 July, reveal an uncoupling between the 3D genome organisation – also called chromatin topology – and gene expression.
Our chromosomes are compartmentalised into domains. Regulatory regions of the DNA that control gene expression – so-called enhancers – are often located in the same chromatin domains (called topologically associated domains) as their target genes. To date, there are a number of interesting examples where domains restrict the activity of enhancers to only the genes within their domain.
To test the generality of these findings more systematically, researchers from the Furlong and Korbel groups at EMBL took advantage of genetic tools in the fruit fly to study highly rearranged mutant chromosomes. Yad Ghavi-Helm, Aleksander Jankowski, Sascha Meiers and colleagues found that changes in chromatin domains were not predictive of changes in gene expression. This means that besides domains, there must be other mechanisms in place that control the specificity of interactions between enhancers and their target genes.
Shaking the dogma
“These results question the generality of a current dogma in the field, that chromatin domains (TADs) are essential to constrain and restrict enhancer function,” says Eileen Furlong, the EMBL group leader who led the study.
“We were able to show that major changes in the 3D organisation of the genome had surprisingly little effect on the expression of most genes, at least in this biological context. The results indicate that while some genes are affected, many appear resistant to rearrangements in their chromatin domain, and that only a small fraction of genes are sensitive to such changes in their topology.”
Enhancers are not that promiscuous
This raises many interesting questions in the field of chromatin topology, for example: what are these other mechanisms that control the interactions between enhancers and their target genes? Many enhancers do not appear to be promiscuous: they do not link to just any target gene, but rather have preferred partners. The team will continue to dissect this by using genetics, optogenetics (a technique to control protein activity with laser light) and single-cell approaches. This will allow them to study the impact of many more perturbations to chromatin topology in both cis and trans.