The closest look ever at the cell’s machines
The first genome-wide screen for protein complexes is completed
Today researchers in Germany announce they have finished the first complete analysis of the “molecular machines” in one of biology’s most important model organisms: S. cerevisiae (baker’s yeast). The study from the biotechnology company Cellzome, in collaboration with the European Molecular Biology Laboratory (EMBL), appears in this week’s online edition of Nature.
“To carry out their tasks, most proteins work in dynamic complexes that may contain dozens of molecules,” says Giulio Superti-Furga, who launched the large-scale project at Cellzome four years ago. “If you think of the cell as a factory floor, up to now, we’ve known some of the components of a fraction of the machines. That has seriously limited what we know about how cells work. This study gives us a nearly complete parts list of all the machines, and it goes beyond that to tell us how they populate the cell and partition tasks among themselves.” The study combined a method of extracting complete protein complexes from cells (tandem affinity purification, developed in 2001 by Bertrand Séraphin at EMBL), mass spectrometry and bioinformatics to investigate the entire protein household of yeast, turning up 257 machines that had never been observed. It also revealed new components of nearly every complex already known.
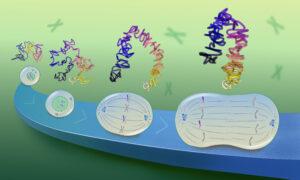
In the course of the work, new computational techniques were developed at EMBL that gave new insights into the dynamic nature of protein complexes. In contrast to most man-made factories, cells continually dismantle and reassemble their machines at different stages of the cellcycle and in response to environmental challenges, such as infections.
“This would be a logistical nightmare if the cell had to build every machine from scratch any time it needed to do something,” says Anne-Claude Gavin, former Director of Molecular and Cell Biology at Cellzome and currently a team leader at EMBL. “We’ve discovered that the reality is different. Cells use a mixed strategy of prefabricating core elements of machines and then synthesizing additional, snap-on molecules that give each machine a precise function. That provides an economic way to diversify biological processes and also to control them.”
Thus if the cell needs to respond quickly, such as in a disease or another emergency, it may only need to produce few parts to switch on or tune the machine. On the other hand, if something shouldn’t happen, it may only need to block the production of a few molecules.
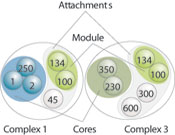
Patrick Aloy and Rob Russell at EMBL used sophisticated computer techniques to reveal the modular organisation of these cellular machines. “This is the most complete set of protein complexes available and probably the set with the highest quality,” Aloy says. “Most proteomics studies in the past have shown whether molecules interact or not, in a ‘yes/no’ way. The completeness of this data lets us see how likely any particular molecule is to bind to another. By combining such measurements for all the proteins in the cell, we discovered new complexes and revealed their modular nature.”
“Investigating protein complexes has always posed a tricky problem – they’re too small to be studied by microscopes, and generally too large to be studied b ytechniques like X-ray crystallography,” says Russell. “But they play such a crucial role in the cell that we need to fill in this gap. There’s still a huge amount to be learned from this data and from the methods we are developing to combine computational and biochemical investigations of the cell.”
“This is an important milestone towards a more global and systems-wide understanding of the cells of organisms ranging from yeast to humans,” says Peer Bork, Head of the Structural and Computational Biology Unit at EMBL, and one of the authors of the paper. “Ultimately we hope to achieve a ‘molecular anatomy’ that takes us from the level of the entire cell to the much deeper level of all the molecules and atoms that make it up.”
Baker’s yeast is evolutionary related to the cells of animals and humans, which means that the findings will be more widely applicable. “The same principles discovered here in yeast apply to human cells,” says Gitte Neubauer, Vice President at Cellzome. “Drug targets and pathologically relevant proteins are parts of machines and pathways.”
The collaboration between Cellzome and the EMBL has been very successful, she says, producing fundamental new insights in how molecules are organised and contributing to Cellzome’s success in complex and pathway analysis.